Research Articles
Mitigating DNA Extraction Kit Batch Effects: A Comprehensive Guide for Robust Genomic Research
This article provides a systematic framework for researchers, scientists, and drug development professionals to identify, understand, and mitigate batch effects introduced by commercial DNA extraction kits.
Unlocking Hidden Microbial Worlds: Advanced Strategies for DNA Extraction from Dormant Microorganisms
This article provides a comprehensive guide for researchers and industry professionals on extracting high-quality DNA from dormant microbial states.
Mastering Variability in Research: How DNA Extraction Methods Drive Experimental Outcomes
This article examines DNA extraction as the primary, often underestimated, source of experimental variability in biomedical research.
Developing a Robust DIY Stool Collection Kit: A Comprehensive Protocol for Reproducible Human Microbiome Research
This article provides a detailed, step-by-step protocol for developing and implementing a standardized do-it-yourself (DIY) stool collection kit for human microbiome studies, modeled on the rigor of the Human Microbiome...
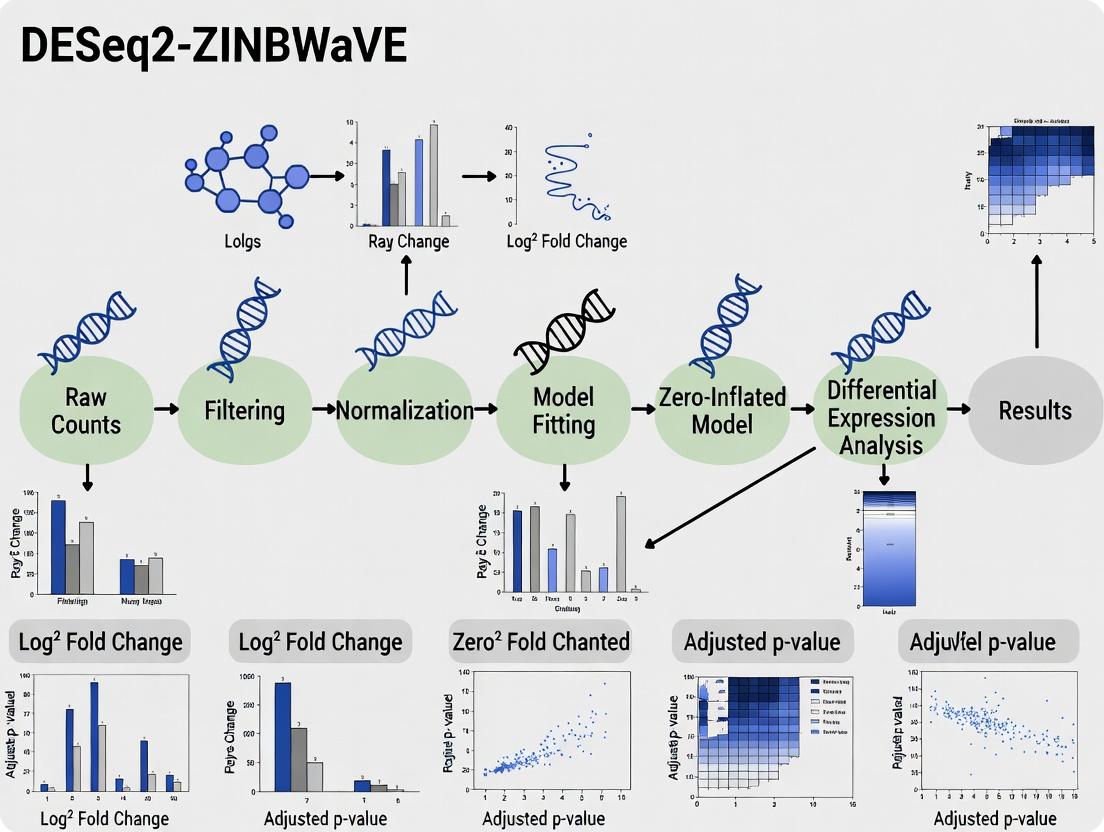
DESeq2-ZINBWaVE Pipeline: A Comprehensive Guide to Differential Abundance Analysis for Biomedical Research
This article provides a complete roadmap for implementing the DESeq2-ZINBWaVE pipeline for differential abundance analysis of high-throughput sequencing data, particularly suited for sparse data like single-cell RNA-seq or microbiome data.
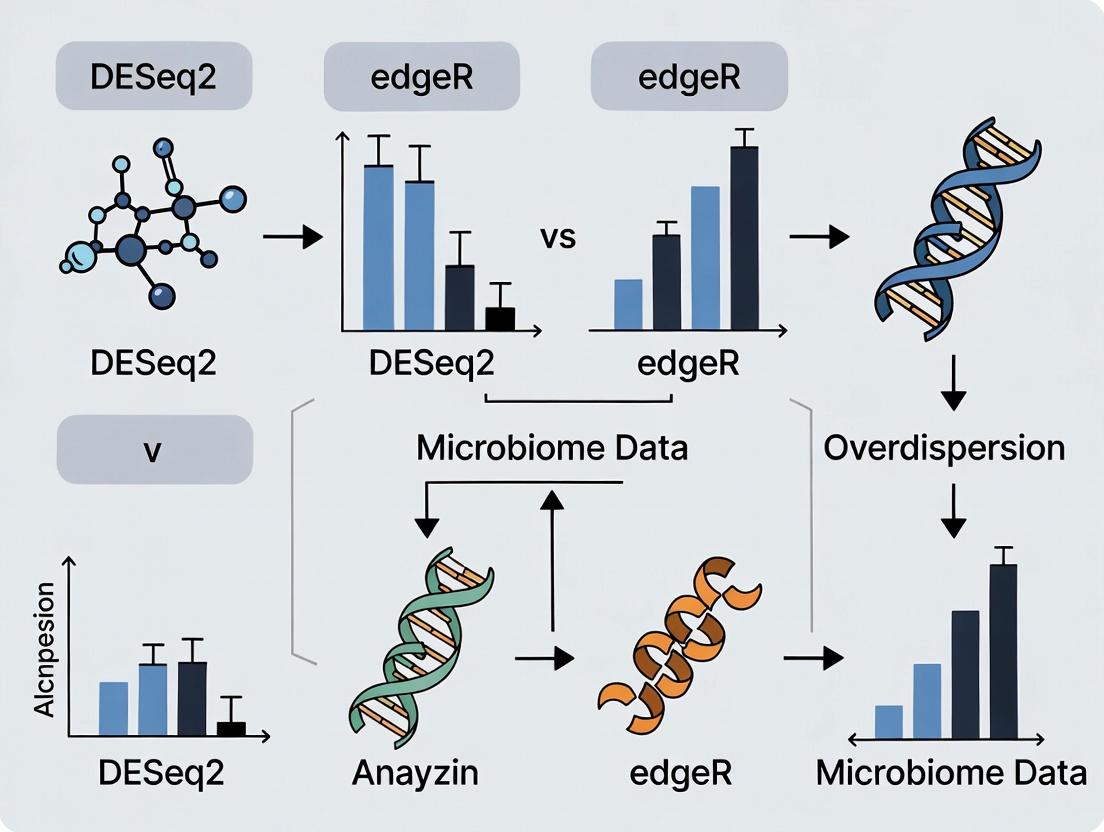
DESeq2 vs edgeR for Microbiome Data: A 2024 Guide for Robust Differential Abundance Analysis
This article provides a comprehensive, practical comparison of DESeq2 and edgeR for analyzing overdispersed microbiome count data.
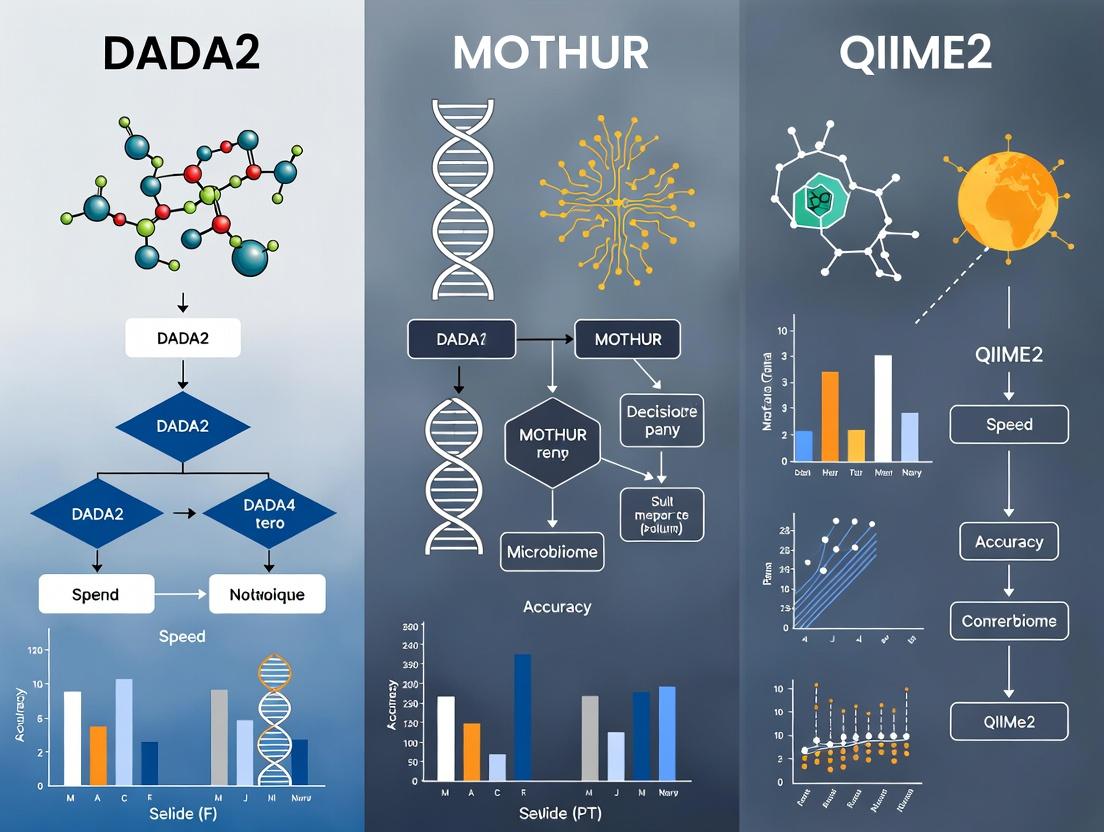
DADA2 vs MOTHUR vs QIIME2: A 2024 Performance Comparison for Microbiome Research and Drug Development
This comprehensive guide analyzes the performance, methodology, and optimal application of the three dominant 16S rRNA analysis pipelines—DADA2, MOTHUR, and QIIME2.
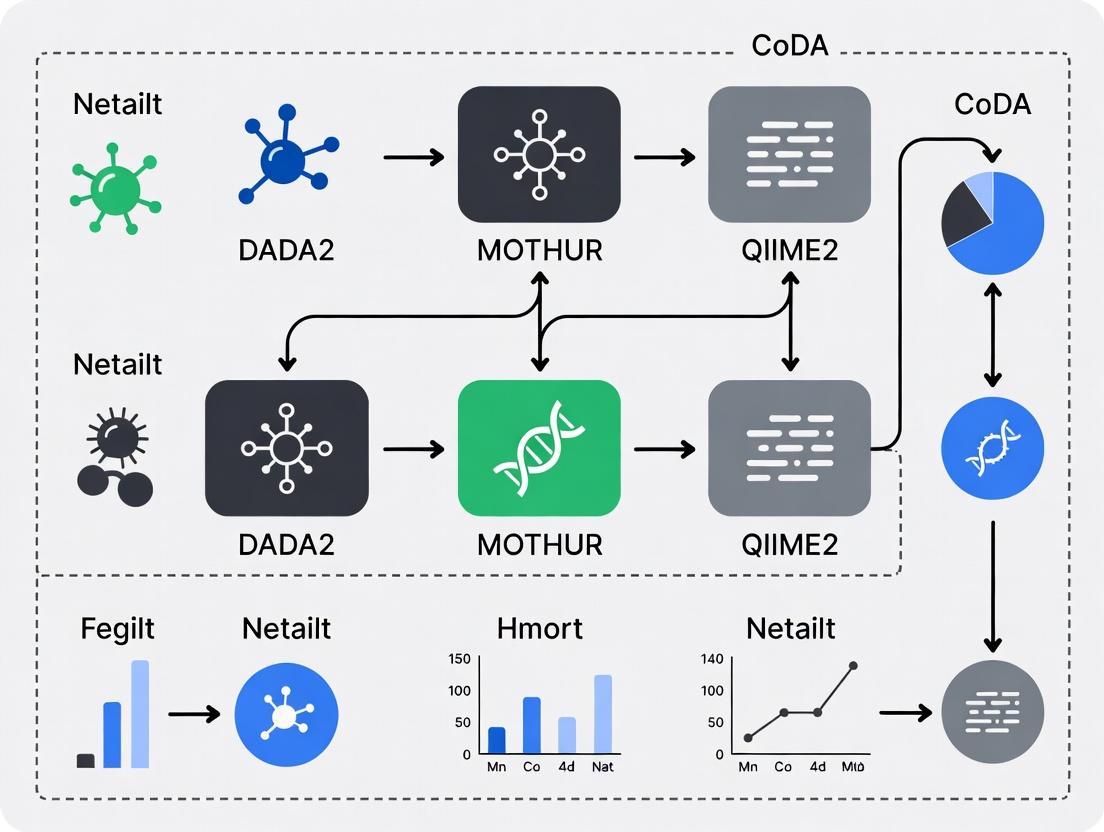
Compositional Data Analysis in Microbiome Research: A Comparative Performance Benchmark of DADA2, MOTHUR, and QIIME2
This article provides a comprehensive, data-driven evaluation of how DADA2, MOTHUR, and QIIME2 perform when their outputs are subjected to Compositional Data Analysis (CoDA) in biomedical research contexts.
Mastering DADA2 Truncation: A Complete Guide to Quality Score Thresholds for Accurate Amplicon Sequence Variants
This comprehensive guide details the critical role of quality score thresholds in the DADA2 pipeline's truncation step for 16S rRNA and other amplicon sequencing data.
Mastering DADA2 Truncation: A Comprehensive Guide to Optimal truncLen-F and truncLen-R Settings for Amplicon Analysis
This article provides researchers, scientists, and drug development professionals with a definitive guide to the DADA2 `truncLen-f` and `truncLen-r` parameters.