Research Articles
Optimizing DNA Extraction for Stool Microbiome Analysis: A Comprehensive Guide for Biomedical Researchers
This article provides a detailed, up-to-date guide on DNA extraction methods for stool microbiome analysis, tailored for researchers, scientists, and drug development professionals.
DADA2 vs MOTHUR vs QIIME2: A 2024 Performance Comparison for Microbiome Research and Drug Development
This comprehensive guide analyzes the performance, methodology, and optimal application of the three dominant 16S rRNA analysis pipelines—DADA2, MOTHUR, and QIIME2.
Compositional Data Analysis in Microbiome Research: A Comparative Performance Benchmark of DADA2, MOTHUR, and QIIME2
This article provides a comprehensive, data-driven evaluation of how DADA2, MOTHUR, and QIIME2 perform when their outputs are subjected to Compositional Data Analysis (CoDA) in biomedical research contexts.
Mastering DADA2 Truncation: A Complete Guide to Quality Score Thresholds for Accurate Amplicon Sequence Variants
This comprehensive guide details the critical role of quality score thresholds in the DADA2 pipeline's truncation step for 16S rRNA and other amplicon sequencing data.
Mastering DADA2 Truncation: A Comprehensive Guide to Optimal truncLen-F and truncLen-R Settings for Amplicon Analysis
This article provides researchers, scientists, and drug development professionals with a definitive guide to the DADA2 `truncLen-f` and `truncLen-r` parameters.
Mastering DADA2 Quality Plots: A Complete Guide to Truncation Parameter Selection for Accurate Amplicon Sequencing Analysis
This comprehensive guide empowers researchers and bioinformaticians to expertly interpret DADA2 quality score plots and determine optimal read truncation parameters.
A Complete Guide to the DADA2 Pipeline for 16S rRNA Analysis: From Raw Reads to ASV Tables
This comprehensive tutorial provides researchers, scientists, and drug development professionals with a complete workflow for analyzing 16S rRNA amplicon sequencing data using the DADA2 pipeline in R.
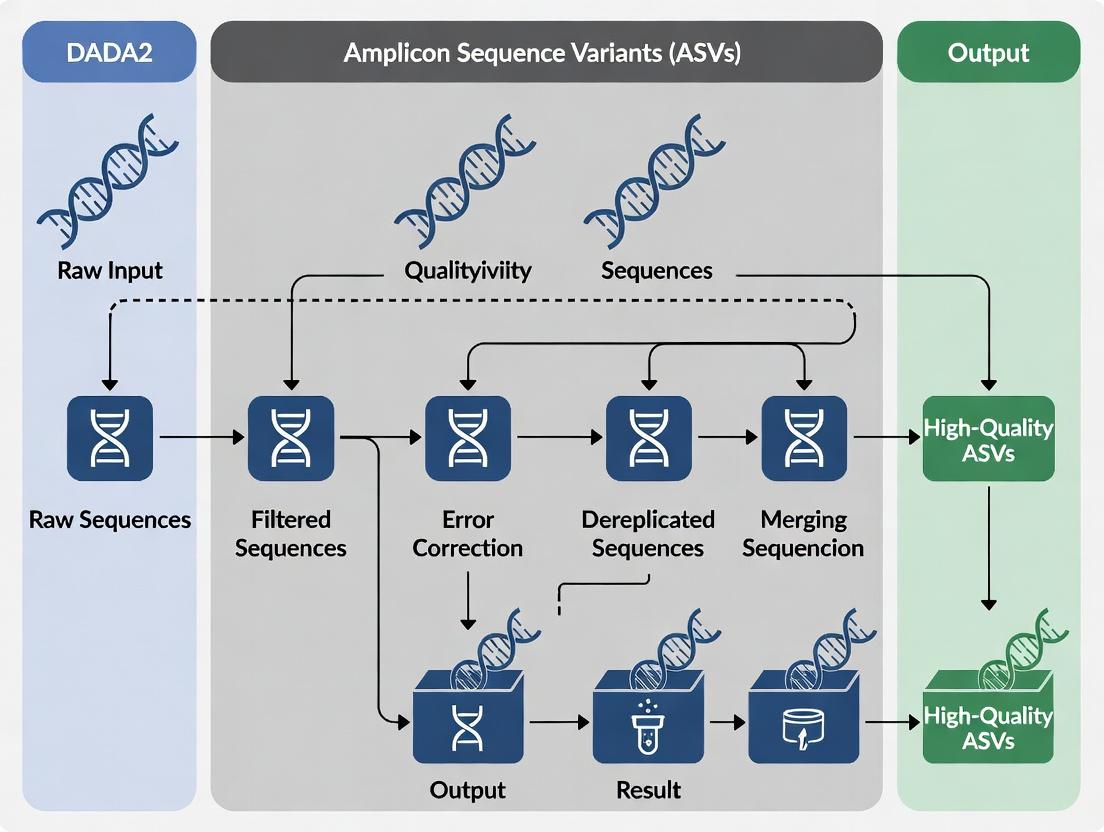
DADA2 ASV Pipeline: A Comprehensive Guide for Accurate Microbiome Analysis in Biomedical Research
This article provides a detailed, practical guide to the DADA2 pipeline for generating high-resolution Amplicon Sequence Variants (ASVs) from 16S rRNA gene sequencing data.
DADA2 Pipeline for ASVs: A Comprehensive Guide for Biomedical Research and Microbiome Analysis
This article provides a complete guide to the DADA2 (Divisive Amplicon Denoising Algorithm) pipeline for generating Amplicon Sequence Variants (ASVs), tailored for researchers and professionals in microbiology, drug development, and...
From Raw Reads to Biological Insights: A Comprehensive DADA2 Pipeline Tutorial for 16S rRNA Amplicon Analysis
This guide provides a complete, step-by-step framework for implementing the DADA2 pipeline to analyze 16S rRNA gene amplicon sequencing data.